 Run Analysis
Documentation
Contact us
Run Analysis
Documentation
Contact us

OpenCustomDB is an innovative tool to build personalized proteogenomic databases. OpenCustomDB uses sample-specific DNA or RNAseq data to acknowledge genomic variants and uses reference or deep genome annotations to generate custom databases of protein sequences.
Regarding deep genome annotation, OpenCustomDB uses OpenProt and OpenVar to ease the detection of unannotated proteins and biomarkers.
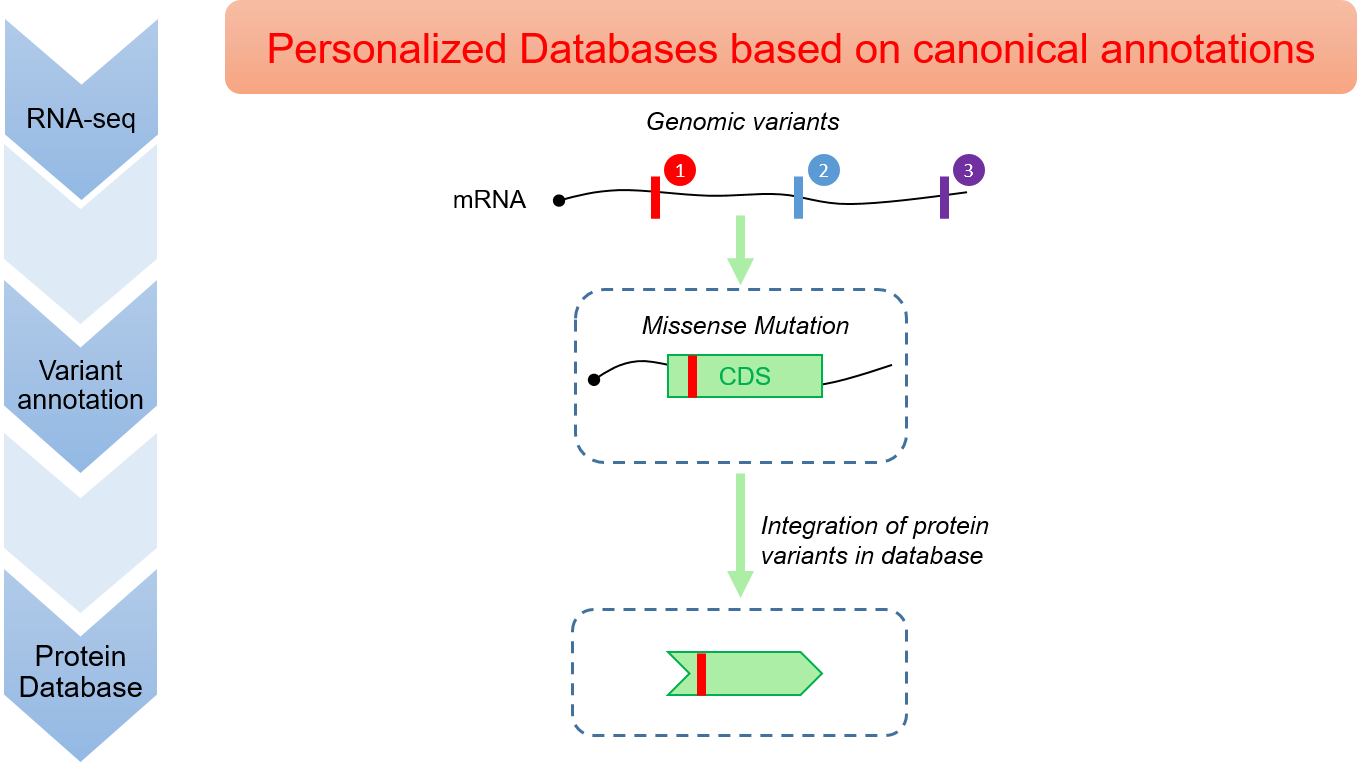
As precision medicine develops, comes the need for proteomics methods that integrate genetic variants to build sample-specific protein databases. However, common methods rely on reference genome annotations that are incomplete. Here, variant #1 yields a missense mutation, variant #2 yields a synonymous mutation, and variant #3 falls outside of any annotated CDS.
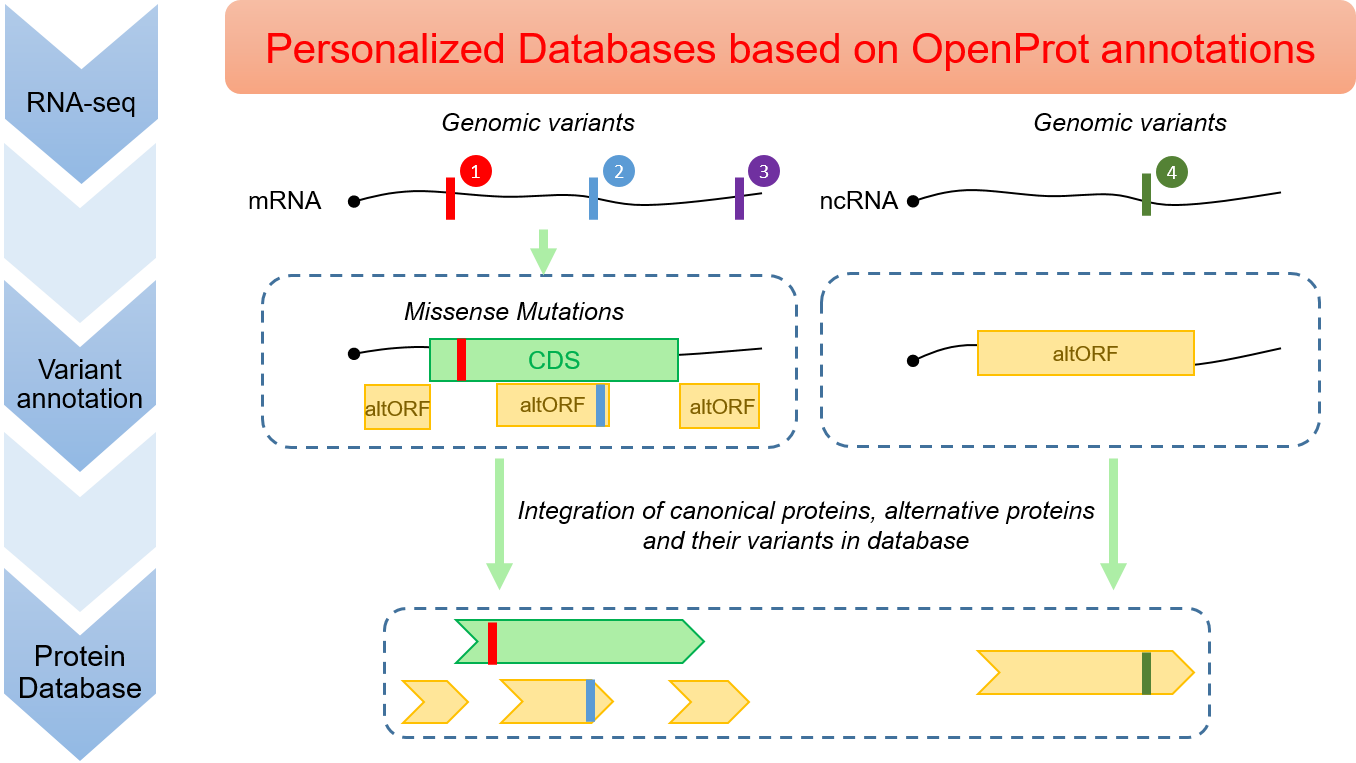
OpenCustomDB uses the OpenProt annotation to build personalised protein databases for proteomics. It enables the detection of alternative proteins and their variants, in addition to canonical proteins and their variants. Here, variant #1 yields a missense mutation; variant #2 yields a synonymous mutation in the canonical CDS and a missense mutation in the shifted reading frame of the altORF. Variant #3 falls outside of any canonical CDS and yields a synonymous mutation in the altORF. Variant #4 yields a missense mutation in the altORF. Alternative proteins and their variants cannot be detected using conventional methods to build personalized databases.